Pyranges is a Python library with a Rust backend for efficient and intuitive manipulation of genomics data, particularly genomic intervals (like genes, genomic features, or reads). The library is optimized for fast querying and manipulation of genomic annotations. It enables intuitive and highly efficient pipelines for genomic analysis.
"Finally ... This was what Python badly needed for years." - Heng Li
This is version 1.x of pyranges. It is a complete rewrite of the original pyranges library, soon to replace the "default" original one (version 0). If you are a v0 user, check the migration guide in the documentation.
The pyranges documentation, including installation instructions, API, tutorial, and how-to-pages, is available at https://pyranges1.readthedocs.io/
# 1.3.8 (21.04.26)
- repo name changed from pyranges_1.x to pyranges1
- updated references to it
- fix issue 151 (Proper use of args and kwargs in concat)
# 1.3.7 (16.04.26)
- require `ruranges>=0.1.4`
- add `preserve_input_order` to Rust-backed overlap-style operations so large results can skip the extra output reordering step
- document the new output-order option with updated docstrings and doctest examples
# 1.3.6 (27.03.26)
- require `ruranges>=0.1.3`
- pick up the `ruranges` fix for `contained_intervals_only=True` overlaps when intervals share the same start coordinate
- fix `merge_overlaps` docs to refer to `count_col` instead of a nonexistent `count` parameter
# 1.3.5 (18.03.26)
- move GTF reading and attribute parsing onto the new `gtfreader>=0.2.0` dependency
- preserve semicolons inside quoted GTF attributes and keep duplicate quoted attributes in `to_rows_keep_duplicates`
- read GTF `Source` and `Frame` columns as categorical dtypes
- update GTF docs and doctests to reflect the new categorical display and supported duplicate-attribute format
- speed up GTF parsing
# 1.3.4 (14.03.26)
- accept ; in quotes in gtf attrs
- also speed up gtf parsing
# 1.3.3 (13.03.26)
- nearest_ranges: treat touching intervals as nearest matches with distance 1 instead of overlapping matches
- document touching-interval nearest behavior in doctests
# 1.3.2 (27.02.26)
- pandas 3 compatibility: removed pandas<3 constraint and aligned test matrix/dependencies
- update doctests/unit test expectations to pandas 3 native formatting (including `str` dtype display)
- remove dtype display normalization workaround in table rendering
- fix pandas 3 copy-on-write/read-only array issue in coverage path used by bigwig/rle conversion
- improve groupby `prod` compatibility across pandas 2/3 edge cases
Pyranges1 requires python ≥3.12. Minimal installation:
pip install pyranges1This installs and requires ruranges>=0.1.3 automatically.
Installation including all optional dependencies:
pip install pyranges1[all]Details at https://pyranges1.readthedocs.io/en/latest/installation.html
- fast
- memory-efficient
- featureful
- pythonic/pandastic
For v1:
Stovner EB, Ticó M, Muñoz del Campo E, Pallarès-Albanell J, Chawla K, Sætrom P, Mariotti M (2025) Pyranges v1: a Python framework for ultrafast sequence interval operations. bioRxiv 2025.12.11.693639; doi: https://doi.org/10.64898/2025.12.11.693639
For v0:
Stovner EB, Sætrom P (2020) PyRanges: efficient comparison of genomic intervals in Python. Bioinformatics 36(3):918-919 http://dx.doi.org/10.1093/bioinformatics/btz615
- most importantly, cite pyranges if you use it. It is the main metric funding sources care about.
- use pyranges in Stack Overflow/biostars questions and answers
- star the repo (possibly important for github visibility and as a proxy for project popularity)
If you encounter bugs, or the documentation is not enough a cannot accomplish a specific task of interest, or if you'd like new features implemented, open an Issue at github: https://github.com/pyranges/pyranges/issues
Pyranges accepts code contributions in form of pull request. For details, visit https://pyranges1.readthedocs.io/developer_guide.html
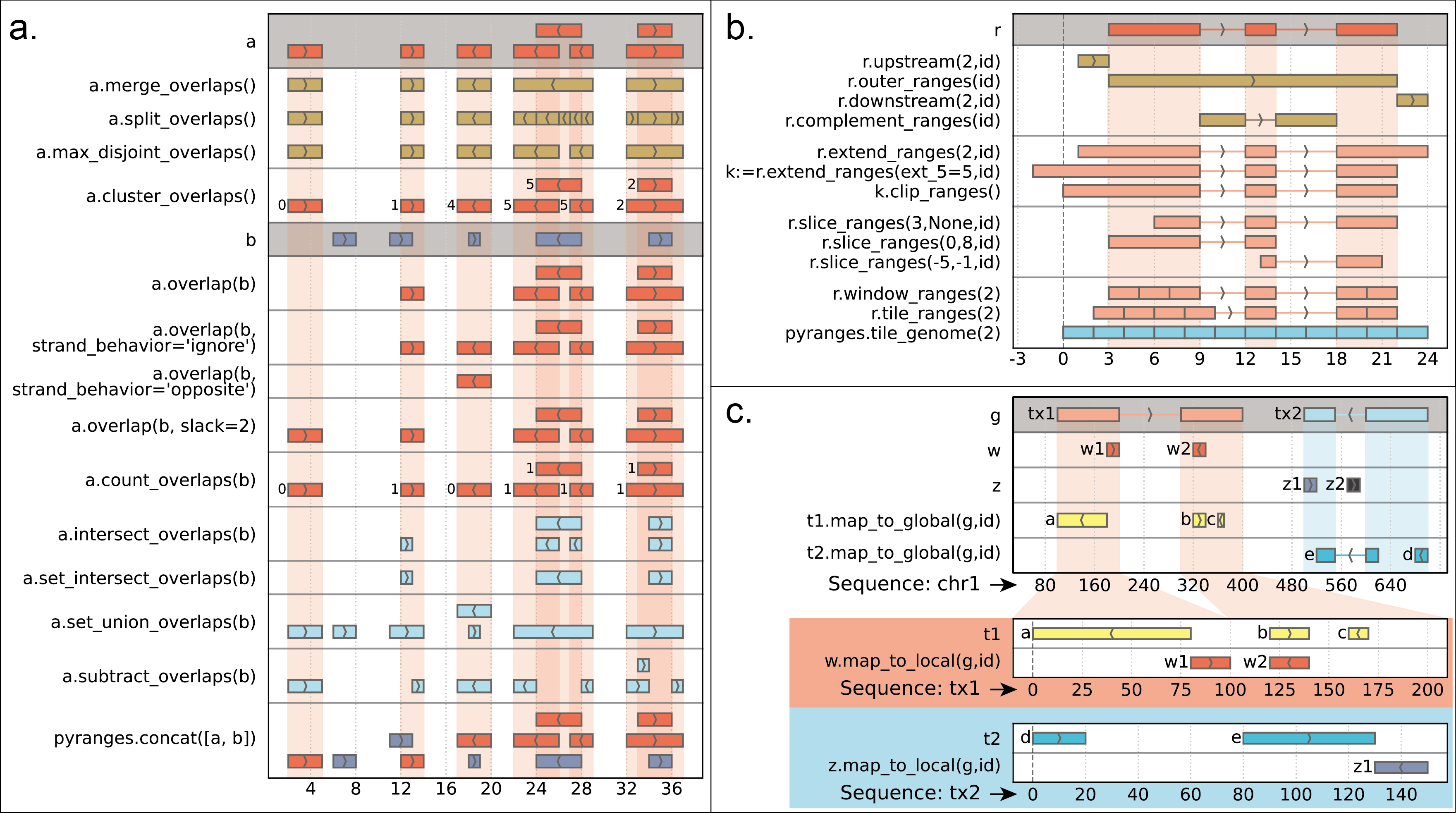 (The cheatsheet above was created with pyrangeyes, a companion graphical library: https://pyrangeyes.readthedocs.io/)
(The cheatsheet above was created with pyrangeyes, a companion graphical library: https://pyrangeyes.readthedocs.io/)